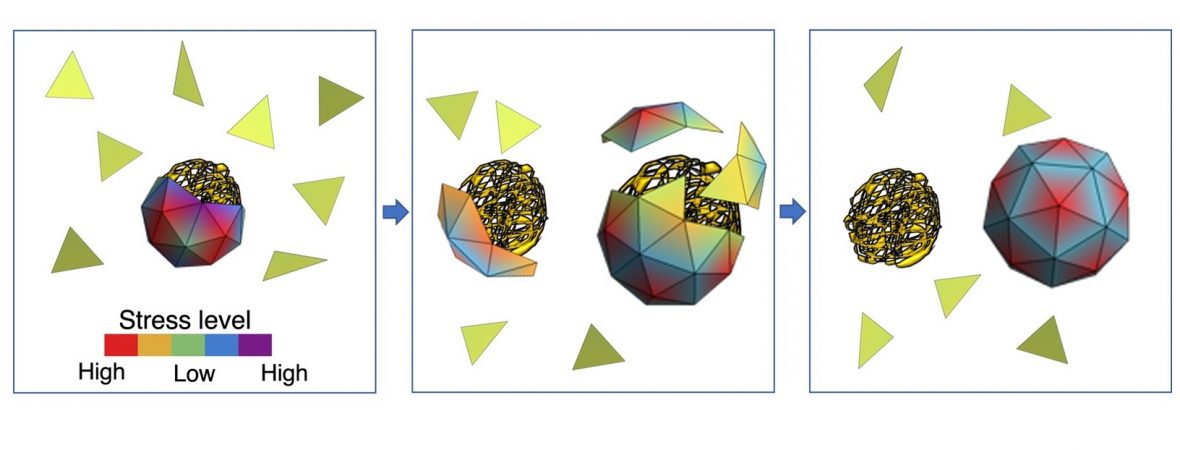
Simulations by UC Riverside-led team could help design nanocontainers used in drug delivery Each simple RNA virus has a genome, its “native RNA.” This genome dictates how the virus replicates in cells to eventually cause disease. The genome also has the code for making a capsid, the protein shell of a virus that encapsulates the genome and protects it like a nanocontainer. A team led by Roya Zandi, a professor of physics and astronomy at the University of California, Riverside, has developed a theory and performed a series of simulations that may help explain how a virus finds its native genome and how capsids form around it and not around other RNAs in the cell. “A better understanding of how capsids form is of vital importance to material scientists and a crucial step in the design of engineered nano-shells that could serve as vehicles for delivering drugs to specific targets in the body,”...
🔒 Premium Content - For Free
Unlock this content by becoming a Global Health Press subscriber. Join for exclusive articles, expert research, and valuable insights!