Rotaviruses, like all viruses, reproduce inside living cells. Making new viruses requires assembling replication factories via a complex, little known process that involves both viral and cellular components. A report in the Proceedings of the National Academy of Sciences by a multidisciplinary team led by researchers at Baylor College of Medicine reveals that the formation of rotavirus factories depends on a cellular protein called CK1α, which chemically modifies viral component NSP2, thus triggering its localization and assembly into the virus factory, an essential step in the formation of new viruses.
“One of the interests of our labs is to better understand the process of assembling rotavirus factories,” said first co-author Dr. Jeanette M. Criglar, staff scientist of molecular virology and microbiology at Baylor College of Medicine and a graduate of the program.
In the process of investigating this, Criglar and her colleagues discovered that a cellular protein called CK1α is required to assemble rotavirus factories.
“When we silenced CK1α in cells before infection with rotavirus, we knocked down the replication of the virus by more than 90 percent, suggesting that CK1α largely controls the formation of rotavirus factories,” Criglar said.
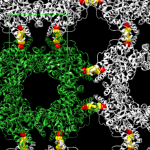
CK1α is a enzyme with the ability to modify other proteins and their functions chemically by adding phosphate groups to them. The researchers discovered that CK1α mediates its effect on the formation of rotavirus replication factories by adding a phosphate group to a rotavirus protein called NSP2. This phosphate modification triggers the assembly of NSP2 octameric units into a crystal-like structure and appears to be required for the formation of rotavirus factories.
“CK1α normally takes care of housekeeping tasks within the cell. Rotavirus takes advantage of this protein’s activity, ‘outsourcing’ it to assemble the virus factories,” said corresponding author Dr. Mary K. Estes, Cullen Foundation Endowed Professor Chair of Human and Molecular Virology at Baylor College of Medicine and emeritus founding director of the Texas Medical Center Digestive Diseases Center.
In addition, the team discovered that rotavirus protein NSP2 can add phosphate groups to itself, thus modifying its activity and affecting other proteins involved in virus assembly. This is a surprising finding, Estes explains, because this function had not been described before for this viral protein.
“Taken together, our findings suggest that a cascade of phosphate chemical modifications, which is mediated in part by CK1α and NSP2, is essential for the formation of rotavirus factories,” said co-author Dr. B V Venkataram Prasad, professor and Alvin Romansky Chair in Biochemistry and Molecular Biology, and member of the Dan L Duncan Comprehensive Cancer Center at Baylor. “These findings provide new insights that could lead to previously unsuspected ways to fight the disease in the future.”
“It is possible that our findings may also shed light on the assembly of virus factories for other viruses that also require CK1α, such as hepatitis C, or that also form cytoplasmic virus factories like West Nile and dengue virus,” Criglar said. “If we can understand how other viruses assemble their factories, perhaps using similar mechanisms to rotavirus, we could advance the understanding of those diseases as well.”
Other authors who contributed to this work include Ramakrishnan Anish (co-first author), Liya Hu, Sue E. Crawford, all at Baylor, and Banumathi Sankaran at Lawrence Berkeley National Laboratory.
Source: Baylor College of Medicine