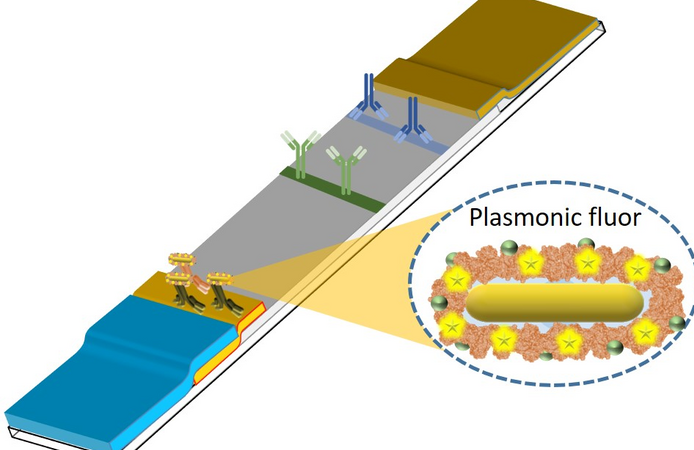
When Srikanth Singamaneni and Guy Genin, both professors of mechanical engineering and materials science at the McKelvey School of Engineering at Washington University in St. Louis, established a new collaboration with researchers from the School of Medicine in late 2019, they didn’t know the landscape of infectious disease research was about to shift dramatically. In a conference room overlooking Forest Park on a beautiful fall day, the team had one goal in mind: tackle the biggest infectious disease problem facing the world right then. “Srikanth and I had a vision of a simple, quantitative diagnostic tool, so we connected with infectious disease physicians here at WashU and asked them, ‘What are the most important questions that could be answered if you could get really detailed information cheaply at the point of care?’” said Genin, the Harold and Kathleen Faught Professor of Mechanical Engineering. “Greg Storch told us that one of the...
🔒 Premium Content - For Free
Unlock this content by becoming a Global Health Press subscriber. Join for exclusive articles, expert research, and valuable insights!