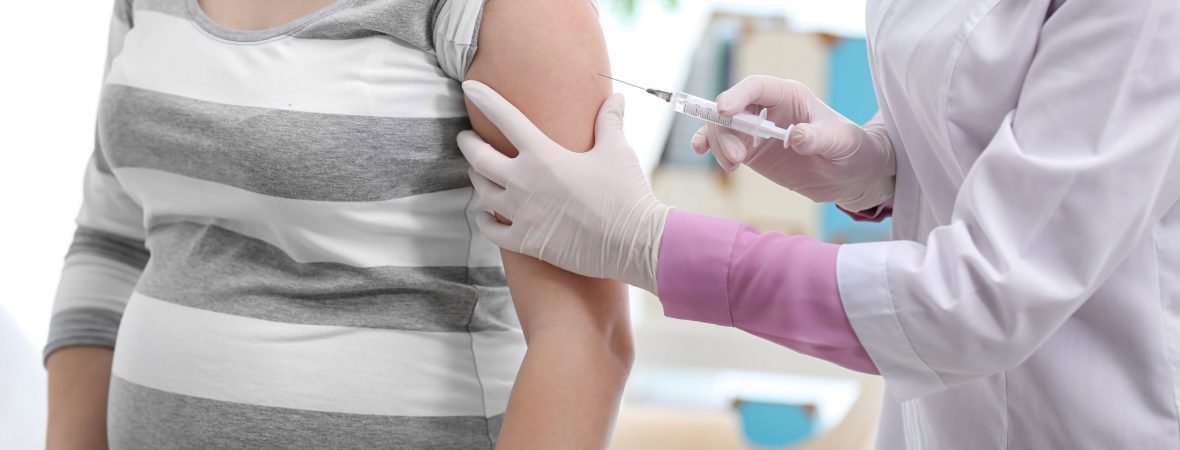
Three genes that could predict susceptibility to congenital Zika virus (ZIKV) infection have been identified, Dr. Irene Rivero-Calle, MD, shared at the annual meeting of the European Society for Paediatric Infectious Diseases, held virtually this year. ZIKV, an emerging flavivirus, is responsible for one of the most critical pandemic emergencies of the last decade and has been associated with severe neonatal brain disabilities, declared Dr. Rivero-Calle, of the Hospital Clínico Universitario de Santiago de Compostela in Santiago de Compostela, Spain. “We think that understanding the genomic background could explain some of the most relevant symptoms of congenital Zika syndrome (CZS) and could be essential to better comprehend this disease.” To achieve this understanding, Dr. Rivero-Calle and her colleagues conducted a study aiming to analyze any genetic factors that could explain the variation in phenotypes in newborns from mothers who had a Zika infection during their pregnancy. Additionally, they strove to “elucidate if...
🔒 Premium Content - For Free
Unlock this content by becoming a Global Health Press subscriber. Join for exclusive articles, expert research, and valuable insights!